CS-466/566: Math for AI
Module 05: Singular Value Decomposition
The University of Alabama
2026-03-23
TABLE OF CONTENTS
Eigen Decomposition: The “Sum of Patterns” View
General diagonalization: \(A = C D C^{-1}\).
For symmetric \(A\): eigenvectors are
orthonormal (\(C^{-1} = C^T\)), so \(A = C D C^T\).
Matrix Multiplication \(\rightarrow\) Sum of Rank-1 Matrices: \[ A = \underbrace{\begin{bmatrix} | & & | \\ v_1 & \dots & v_n \\ | & & | \end{bmatrix}}_{V} \underbrace{\begin{bmatrix} \lambda_1 & & \\ & \ddots & \\ & & \lambda_n \end{bmatrix}}_{\Lambda} \underbrace{\begin{bmatrix} - v_1^T - \\ \vdots \\ - v_n^T - \end{bmatrix}}_{V^T} = \sum_{i=1}^n \underbrace{\lambda_i}_{\text{scalar}} \cdot \underbrace{\mathbf{v}_i}_{n \times 1} \cdot \underbrace{\mathbf{v}_i^T}_{1 \times n} \]
Insight: Each term \(\lambda_i \mathbf{v}_i \mathbf{v}_i^T\) is an \(n \times n\) matrix. If \(\lambda_i \approx 0\), that term vanishes \(\rightarrow\) Sparsity / Dimensionality Reduction!
TABLE OF CONTENTS
Eigen Vectors for Square Matrices Requirement
Let’s consider a non-square matrix \(A\): \[ A \mathbf{x} = \begin{bmatrix} 1 & 0 \\ 0 & 1 \\ 0 & 0 \end{bmatrix}_{(3 \times 2)} \begin{bmatrix} x_1 \\ x_2 \end{bmatrix}_{(2 \times 1)} \]
if \(x \in \mathbb{R}^2\) then \(Ax \in \mathbb{R}^3\)
So it can never equal \(\lambda x \in \mathbb{R}^2\).
Eigen decomposition only works for square matrices!
Why Eigen Decomposition is Not Always Possible
Eigen decomposition / diagonalization assumes:
- \(A\) is square (\(n \times n\))
- and has enough linearly independent eigenvectors
But ML data is usually rectangular
- \(X\) = (number of samples) \(\times\) (number of features)
- e.g., \(X \in \mathbb{R}^{10000 \times 768}\)
So… How to find a similar concept to eigenvectors for a non-square matrix?
This is exactly why we use SVD.
Trick: Build Two Square Matrices from Any \(A\)
Let \(A\) be any \(m \times n\) matrix (maybe not square).
We can always form: \[ AA^T \in \mathbb{R}^{m \times m}, \quad A^TA \in \mathbb{R}^{n \times n} \] Both are:
- square
- symmetric
- eigenvalues are \(\ge 0\)
So even if \(A\) is rectangular, \(AA^T\) and \(A^TA\) have real eigenvectors.
Why is \(AA^T\) Symmetric and Square?
Square: If \(A\) is \(m \times n\), then \(A^T\) is \(n \times m\). \[ \underbrace{A}_{m \times n} \cdot \underbrace{A^T}_{n \times m} = \underbrace{AA^T}_{m \times m} \]
\[ \underbrace{\begin{bmatrix} 1 & 2 \\ 3 & 4 \\ 5 & 6 \end{bmatrix}}_{3 \times 2} \cdot \underbrace{\begin{bmatrix} 1 & 3 & 5 \\ 2 & 4 & 6 \end{bmatrix}}_{2 \times 3} = \underbrace{\begin{bmatrix} 5 & 11 & 17 \\ 11 & 25 & 39 \\ 17 & 39 & 61 \end{bmatrix}}_{3 \times 3} \]
SVD Roadmap
Our Goal: Factor any \(m \times n\) matrix \(A\) into \(A = U \Sigma V^T\).
The 3-Step Journey:
- Find \(V\)?
- Find \(\Sigma\) (Singular values)?
- Find \(U\)?
Since \(A^TA\) is symmetric → we can always find orthogonal eigenvectors in an indirect way!
SVD Derivation (Part 1)
Step 1: Start with \(A^TA\)
We can always find the eigenvalues and eigenvectors of \(A^TA\) (even if \(A\) is not square): \[ \underbrace{A^TA}_{n \times n} \underbrace{\mathbf{v}_i}_{\substack{\text{right eigenvectors} \\ n \times 1}} = \lambda_i \underbrace{\mathbf{v}_i}_{\substack{\text{right eigenvectors} \\ n \times 1}} \] These \(\mathbf{v}_i\) are our Right Singular Vectors.
Step 2: Define Singular Values
Since \(\lambda_i \ge 0\): \(\quad \sigma_i = \sqrt{\lambda_i}\)
\[ \underbrace{A^TA}_{n \times n} \underbrace{\mathbf{v}_i}_{\substack{\text{right eigenvectors} \\ n \times 1}} = \sigma_i^2 \underbrace{\mathbf{v}_i}_{\substack{\text{right eigenvectors} \\ n \times 1}} \]
SVD Derivation (Part 2)
Step 3: Multiply both sides by \(A\): \[ \underbrace{(AA^T)}_{m \times m}\underbrace{(A\mathbf{v}_i)}_{\substack{\text{eigenvector!} \\ m \times 1}} = \sigma_i^2 \underbrace{(A\mathbf{v}_i)}_{\substack{\text{eigenvector!} \\ m \times 1}} \]
So \(A\mathbf{v}_i\) is an eigenvector of \(AA^T\)!
Define: \(\mathbf{u}_i = \frac{A\mathbf{v}_i}{\sigma_i}\) (normalized).
Then: \(A \mathbf{v}_i = \sigma_i \mathbf{u}_i\) ← core SVD relation!
Step 4: Stack & Finish
Stack vectors: \(\underbrace{V}_{n \times n} = [\mathbf{v}_1, \dots, \mathbf{v}_n], \quad \underbrace{U}_{m \times m} = [\mathbf{u}_1, \dots, \mathbf{u}_m]\)
The key relation: \(A\mathbf{v}_i = \sigma_i \mathbf{u}_i \implies AV = U\Sigma \implies\) \(A = U \Sigma V^T\) ✅
The One Relationship to Remember
SVD gives paired vectors \((u_i, v_i)\) such that: \[ \underbrace{A}_{m \times n} \cdot \quad \underbrace{v_i}_{n \times 1} \quad = \quad \sigma_i \quad \underbrace{u_i}_{m \times 1} \] Meaning:
- \(v_i\): a special input direction
- \(A v_i\): lands exactly on direction \(u_i\)
- scaled by \(\sigma_i\)
So \(A\) maps one special axis to another special axis, with a known scale.
Numeric Example (1/8)
\[ A = \begin{bmatrix} 3 & 2 & 2 \\ 2 & 3 & -2 \end{bmatrix}_{2 \times 3} \quad \implies \quad A^T = \begin{bmatrix} 3 & 2 \\ 2 & 3 \\ 2 & -2 \end{bmatrix}_{3 \times 2} \]
Step 1: Compute \(A^T A\): \[ A^T A = \begin{bmatrix} 3 & 2 \\ 2 & 3 \\ 2 & -2 \end{bmatrix}_{3 \times 2} \begin{bmatrix} 3 & 2 & 2 \\ 2 & 3 & -2 \end{bmatrix}_{2 \times 3} = \begin{bmatrix} 13 & 12 & 2 \\ 12 & 13 & -2 \\ 2 & -2 & 8 \end{bmatrix}_{3 \times 3} \]
Numeric Example (2/8)
Step 2: Eigenvalues of \(A^T A\) — solve \(\det(A^T A - \lambda I) = 0\): \[ \det\left( \begin{bmatrix} 13 - \lambda & 12 & 2 \\ 12 & 13 - \lambda & -2 \\ 2 & -2 & 8 - \lambda \end{bmatrix} \right) = 0 \]
Cofactor expansion along row 1: \[ = (13-\lambda) \det\begin{bmatrix} 13-\lambda & -2 \\ -2 & 8-\lambda \end{bmatrix} - 12 \det\begin{bmatrix} 12 & -2 \\ 2 & 8-\lambda \end{bmatrix} + 2 \det\begin{bmatrix} 12 & 13-\lambda \\ 2 & -2 \end{bmatrix} \]
Compute each 2×2 determinant: \[ = (13-\lambda)[(13-\lambda)(8-\lambda) - 4] - 12[12(8-\lambda) + 4] + 2[-24 - 2(13-\lambda)] \]
\[ = \lambda^3 - 34\lambda^2 + 225\lambda = \lambda(\lambda - 25)(\lambda - 9) = 0 \] \(\implies \lambda_1 = 25, \quad \lambda_2 = 9, \quad \lambda_3 = 0\)
Numeric Example (3/8)
Step 3: Singular Values (\(\sigma = \sqrt{\lambda}\)): \[ \sigma_1 = \sqrt{25} = 5, \quad \sigma_2 = \sqrt{9} = 3, \quad \sigma_3 = \sqrt{0} = 0 \]
The non-zero singular values are \(\sigma_1 = 5\) and \(\sigma_2 = 3\).
Since \(A\) is \(2 \times 3\), we have at most 2 non-zero singular values.
Numeric Example (4/8): Find \(V\) (Part a)
Step 4: Find eigenvectors of \(A^TA\) for each \(\lambda\)
For \(\lambda_1 = 25\): solve \((A^TA - 25I)\mathbf{v}_1 = 0\) \[ \begin{bmatrix} 13-25 & 12 & 2 \\ 12 & 13-25 & -2 \\ 2 & -2 & 8-25 \end{bmatrix} \mathbf{v}_1 = \begin{bmatrix} -12 & 12 & 2 \\ 12 & -12 & -2 \\ 2 & -2 & -17 \end{bmatrix} \mathbf{v}_1 = \mathbf{0} \]
Row reduce → from row 1: \(-12v_1 + 12v_2 + 2v_3 = 0 \implies v_1 = v_2 + \frac{v_3}{6}\)
Setting \(v_3 = 0, v_2 = 1 \implies v_1 = 1\). Normalize: \[ \mathbf{v}_1 = \frac{1}{\sqrt{2}}\begin{bmatrix} 1 \\ 1 \\ 0 \end{bmatrix} \]
Numeric Example (5/8): Find \(V\) (Part b)
For \(\lambda_2 = 9\): solve \((A^TA - 9I)\mathbf{v}_2 = 0\) \[ \begin{bmatrix} 13-9 & 12 & 2 \\ 12 & 13-9 & -2 \\ 2 & -2 & 8-9 \end{bmatrix} \mathbf{v}_2 = \begin{bmatrix} 4 & 12 & 2 \\ 12 & 4 & -2 \\ 2 & -2 & -1 \end{bmatrix} \mathbf{v}_2 = \mathbf{0} \]
Row reduce → solution: \(v_1 = 1, v_2 = -1, v_3 = 4\) (proportional). Normalize: \[ \mathbf{v}_2 = \frac{1}{\sqrt{18}}\begin{bmatrix} 1 \\ -1 \\ 4 \end{bmatrix} \]
Numeric Example (6/8): Find \(V\) (Part c)
For \(\lambda_3 = 0\): solve \((A^TA - 0I)\mathbf{v}_3 = 0\) \[ \begin{bmatrix} 13 & 12 & 2 \\ 12 & 13 & -2 \\ 2 & -2 & 8 \end{bmatrix} \mathbf{v}_3 = \mathbf{0} \]
Row reduce → solution: \(v_1 = -2, v_2 = 2, v_3 = 1\) (proportional).
Normalize: \[ \mathbf{v}_3 = \frac{1}{3}\begin{bmatrix} -2 \\ 2 \\ 1 \end{bmatrix} \]
Numeric Example (7/8): Find \(U\)
Step 5: Find \(U\) — use \(\mathbf{u}_i = \frac{A\mathbf{v}_i}{\sigma_i}\)
\(\mathbf{u}_1 = \frac{1}{\sigma_1} A \mathbf{v}_1 = \frac{1}{5} \begin{bmatrix} 3 & 2 & 2 \\ 2 & 3 & -2 \end{bmatrix} \cdot \frac{1}{\sqrt{2}}\begin{bmatrix} 1 \\ 1 \\ 0 \end{bmatrix} = \frac{1}{5\sqrt{2}} \begin{bmatrix} 5 \\ 5 \end{bmatrix} = \frac{1}{\sqrt{2}}\begin{bmatrix} 1 \\ 1 \end{bmatrix}\)
\(\mathbf{u}_2 = \frac{1}{\sigma_2} A \mathbf{v}_2 = \frac{1}{3} \begin{bmatrix} 3 & 2 & 2 \\ 2 & 3 & -2 \end{bmatrix} \cdot \frac{1}{\sqrt{18}}\begin{bmatrix} 1 \\ -1 \\ 4 \end{bmatrix} = \frac{1}{3\sqrt{18}} \begin{bmatrix} 9 \\ -9 \end{bmatrix} = \frac{1}{\sqrt{2}}\begin{bmatrix} 1 \\ -1 \end{bmatrix}\)
Numeric Example (8/8)
Step 6: Build \(\Sigma\): \[ \Sigma = \begin{bmatrix} 5 & 0 & 0 \\ 0 & 3 & 0 \end{bmatrix}_{2 \times 3} \]
Final Decomposition (\(A = U \Sigma V^T\)): \[ \underbrace{A}_{2 \times 3} = \underbrace{U}_{2 \times 2} \cdot \underbrace{\Sigma}_{2 \times 3} \cdot \underbrace{V^T}_{3 \times 3} \]
\[ \implies \begin{bmatrix} 3 & 2 & 2 \\ 2 & 3 & -2 \end{bmatrix} = \underbrace{\begin{bmatrix} \frac{1}{\sqrt{2}} & \frac{1}{\sqrt{2}} \\ \frac{1}{\sqrt{2}} & \frac{-1}{\sqrt{2}} \end{bmatrix}}_{U} \underbrace{\begin{bmatrix} 5 & 0 & 0 \\ 0 & 3 & 0 \end{bmatrix}}_{\Sigma} \underbrace{\begin{bmatrix} \frac{1}{\sqrt{2}} & \frac{1}{\sqrt{2}} & 0 \\ \frac{1}{\sqrt{18}} & \frac{-1}{\sqrt{18}} & \frac{4}{\sqrt{18}} \\ \frac{-2}{3} & \frac{2}{3} & \frac{1}{3} \end{bmatrix}}_{V^T} \]
TABLE OF CONTENTS
The Best Low-Rank Approximation
SVD allows us to represent an image (matrix \(A\)) as a sum of rank-1 patterns: \[ A = \sigma_1 u_1 v_1^T + \sigma_2 u_2 v_2^T + \cdots + \sigma_r u_r v_r^T \]
By keeping only the first \(k\) terms, we get the best possible approximation of rank \(k\).

Original

k=5 (Blurry)

k=20 (Better)

k=50 (Almost Perfect)
TABLE OF CONTENTS
SVD in NumPy
In Python, we use numpy.linalg.svd:
Key Details:
Sis returned as a 1D array (vector), not a diagonal matrix.Vtis already transposed (\(V^T\)).full_matrices=Falsecomputes the Economy SVD (faster).
The singular values in \(S\) are returned sorted in descending order: \[ \sigma_1 \ge \sigma_2 \ge \sigma_3 \ge \dots \ge 0 \]
Reconstructing from SVD
To rebuild the matrix (or an approximation), we need to turn S back into a diagonal matrix:
Setting the value of \(k\) controls the rank of the approximation: \[ A_k = U \cdot \Sigma_k \cdot V^T \]
TABLE OF CONTENTS
The Curse of Dimensionality
Real-world data has many features:
- Student records: Height, Weight, GPA, SAT, …
- Images: thousands of pixels
- Text: thousands of word frequencies
- Genomics: ~20,000 genes
As dimensions \(\uparrow\), data becomes sparse, distances become meaningless, and models overfit.

Question: Can we find a smaller set of variables that captures most of the information?
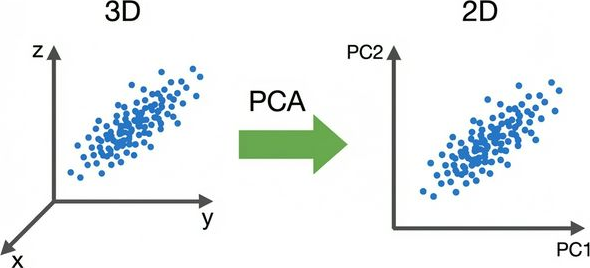
What Does PCA Do?
PCA finds a new coordinate system where:
- Axes point along the directions of maximum variance
- Axes are orthogonal to each other
- We can drop axes with low variance → dimensionality reduction
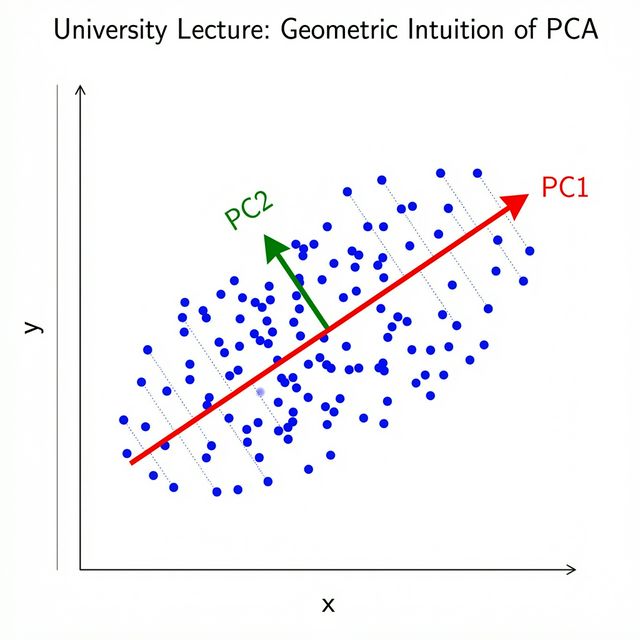
Geometric Intuition
Given a cloud of data points:
- PC1 = direction of greatest spread (variance)
- PC2 = direction of greatest spread perpendicular to PC1
- PC3 = … and so on
Each Principal Component captures the next most important pattern in the data.
PCA finds the best linear summary of your data in fewer dimensions.
PCA Algorithm Summary
| Step | Operation | Result |
|---|---|---|
| 1 | Center: \(\bar{X} = X - \mathbf{1}\boldsymbol{\mu}^T\) | Zero-mean data |
| 2 | SVD: \(\bar{X} = U \Sigma V^T\) | Get components |
| 3 | Choose \(k\) components | Keep top \(k\) columns of \(V\) |
| 4 | Project: \(Z = \bar{X} V_k\) | Reduced data (\(n \times k\)) |
Result: We go from \(d\) features → \(k\) features, where \(k \ll d\).
PCA in Python
Why Does SVD Give Us the Principal Components?
Step 1: Center the data
Given data matrix \(X\) (\(n\) samples \(\times\) \(d\) features), subtract the mean: \[ \bar{X} = X - \mathbf{1}\boldsymbol{\mu}^T \]
Step 2: The covariance matrix is \[ C = \frac{1}{n-1} \bar{X}^T \bar{X} \] Its eigenvectors = principal components, eigenvalues = variance along each PC.
Step 3: Apply SVD
\(\bar{X} = U \Sigma V^T \implies\) the columns of \(V\) are the principal components!
\[ C = \frac{1}{n-1} V \Sigma^2 V^T \]
How Many Components to Keep?
The variance explained by each component is:
\[ \text{Var explained by PC}_i = \frac{\sigma_i^2}{\sum_{j=1}^{d} \sigma_j^2} \]
Rule of thumb: keep enough components to explain 90-95% of total variance.
Example: If \(\sigma_1 = 10, \sigma_2 = 5, \sigma_3 = 1, \sigma_4 = 0.1\)
| PC | \(\sigma_i^2\) | % Variance | Cumulative |
|---|---|---|---|
| 1 | 100 | 79.4% | 79.4% |
| 2 | 25 | 19.8% | 99.2% |
| 3 | 1 | 0.8% | 100.0% |
| 4 | 0.01 | ~0% | 100.0% |
→ Keeping just 2 PCs captures 99.2% of the information!
Thank You!




The University of Alabama